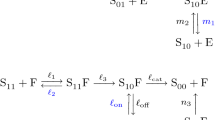Abstract
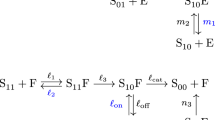
This work investigates the emergence of oscillations in one of the simplest cellular signaling networks exhibiting oscillations, namely the dual-site phosphorylation and dephosphorylation network (futile cycle), in which the mechanism for phosphorylation is processive, while the one for dephosphorylation is distributive (or vice versa). The fact that this network yields oscillations was shown recently by Suwanmajo and Krishnan. Our results, which significantly extend their analyses, are as follows. First, in the three-dimensional space of total amounts, the border between systems with a stable versus unstable steady state is a surface defined by the vanishing of a single Hurwitz determinant. Second, this surface consists generically of simple Hopf bifurcations. Next, simulations suggest that when the steady state is unstable, oscillations are the norm. Finally, the emergence of oscillations via a Hopf bifurcation is enabled by the catalytic and association constants of the distributive part of the mechanism; if these rate constants satisfy two inequalities, then the system generically admits a Hopf bifurcation. Our proofs are enabled by the Routh–Hurwitz criterion, a Hopf bifurcation criterion due to Yang, and a monomial parametrization of steady states.





Similar content being viewed by others
Notes
This file and others mentioned below are in the Supporting Information; see “Appendix A”.
The functions are provided as a text file in the Supporting Information. See “Appendix A”.
References
Aoki K, Yamada M, Kunida K, Yasuda S, Matsuda M (2011) Processive phosphorylation of ERK MAP kinase in mammalian cells. Proc Natl Acad Sci USA 108(31):12675–12680
Atkins P, De Paula J, Keeler J (2018) Atkins’ physical chemistry. Oxford University Press, Oxford
Bure EG, Rozenvasser YN (1974) On investigations of autooscillating system sensitivity, no 7. Avtomat. i Telemekh, pp 9–17
Conradi C, Mincheva M (2014) Catalytic constants enable the emergence of bistability in dual phosphorylation. J R Soc Interface 11(95):20140158
Conradi C, Shiu A (2015) A global convergence result for processive multisite phosphorylation systems. Bull Math Biol 77(1):126–155
Conradi C, Shiu A (2018) Dynamics of post-translational modification systems: recent progress and future challenges. Biophys J 114(3):507–515
Conradi C, Feliu E, Mincheva M, Wiuf C (2017) Identifying parameter regions for multistationarity. PLoS Comput Biol 13(10):e1005751
Dhooge A, Govaerts W, Kuznetsov YA (2003) MATCONT: a MATLAB package for numerical bifurcation analysis of ODEs. ACM Trans Math Softw 29(2):141–164
Domijan M, Kirkilionis M (2009) Bistability and oscillations in chemical reaction networks. J Math Biol 59(4):467–501
Eithun M, Shiu A (2017) An all-encompassing global convergence result for processive multisite phosphorylation systems. Math Biosci 291:1–9
Errami H, Eiswirth M, Grigoriev D, Seiler WM, Sturm T, Weber A (2015) Detection of Hopf bifurcations in chemical reaction networks using convex coordinates. J Comput Phys 291:279–302
Ferrell JE, Ha SH (2014) Ultrasensitivity part II: multisite phosphorylation, stoichiometric inhibitors, and positive feedback. Trends Biochem Sci 39(11):556–569
Gantmacher FR (1959) Matrix theory, vol 21. Chelsea, New York
Gatermann K, Eiswirth M, Sensse A (2005) Toric ideals and graph theory to analyze Hopf bifurcations in mass action systems. J Symb Comput 40(6):1361–1382
Gelfand IM, Kapranov MM, Zelevinsky AV (1994) Discriminants, resultants and multidimensional determinants. Birkhäuser, Basel
Guckenheimer J, Holmes P (2013) Nonlinear oscillations, dynamical systems, and bifurcations of vector fields, vol 42. Springer, Berlin
Hadač O, Muzika F, Nevoral V, Přibyl M, Schreiber I (2017) Minimal oscillating subnetwork in the Huang-Ferrell model of the MAPK cascade. PLoS ONE 12(6):1–25
Hell J, Rendall AD (2015) A proof of bistability for the dual futile cycle. Nonlinear Anal Real 24:175–189
Hilioti Z, Sabbagh W, Paliwal S, Bergmann A, Goncalves MD, Bardwell L, Levchenko A (2008) Oscillatory phosphorylation of yeast Fus3 MAP kinase controls periodic gene expression and morphogenesis. Curr Biol 18(21):1700–1706
Hu H, Goltsov A, Bown JL, Sims AH, Langdon SP, Harrison DJ, Faratian D (2013) Feedforward and feedback regulation of the MAPK and PI3K oscillatory circuit in breast cancer. Cell Signal 25(1):26–32
Ingalls BP (2004) Autonomously oscillating biochemical systems: parametric sensitivity of extrema and period. Syst Biol 1(1):62–70
Ingalls B, Mincheva M, Roussel MR (2017) Parametric sensitivity analysis of oscillatory delay systems with an application to gene regulation. Bull Math Biol 79(7):1539–1563
Johnston MD (2014) Translated chemical reaction networks. Bull Math Biol 76(6):1081–1116
Johnston MD, Müller S, Pantea C (2018) A deficiency-based approach to parametrizing positive equilibria of biochemical reaction systems. Preprint. arXiv:1805.09295
Liu WM (1994) Criterion of Hopf bifurcations without using eigenvalues. J Math Anal Appl 182(1):250–256
Lozada-Cruz G (2012) The simple application of the implicit function theorem, vol XIX, no 1. Boletin de la Asociatión Matemática Venezolana
Millán MP, Dickenstein A (2018) The structure of MESSI biological systems. SIAM J Appl Dyn Syst 17(2):1650–1682
Millán MP, Turjanski AG (2015) MAPK’s networks and their capacity for multistationarity due to toric steady states. Math Biosci 262:125–137
Millán MP, Dickenstein A, Shiu A, Conradi C (2012) Chemical reaction systems with toric steady states. Bull Math Biol 74(5):1027–1065
Müller S, Feliu E, Regensburger G, Conradi C, Shiu A, Dickenstein A (2016) Sign conditions for injectivity of generalized polynomial maps with applications to chemical reaction networks and real algebraic geometry. Found Comput Math 16(1):69–97
Ode KL, Ueda HR (2017) Design principles of phosphorylation-dependent timekeeping in eukaryotic circadian clocks. Cold Spring Harb Perspect Biol 10:a028357
Rao S (2017) Global stability of a class of futile cycles. J Math Biol 74:709–726
Rao S (2018) Stability analysis of the Michaelis-Menten approximation of a mixed mechanism of a phosphorylation system. Math Biosci 301:159–166
Rubinstein BY, Mattingly HH, Berezhkovskii AM, Shvartsman SY (2016) Long-term dynamics of multisite phosphorylation. Mol Biol Cell 27(14):2331–2340
Salazar C, Höfer T (2009) Multisite protein phosphorylation—from molecular mechanisms to kinetic models. FEBS J 276(12):3177–3198
Suwanmajo T, Krishnan J (2015) Mixed mechanisms of multi-site phosphorylation. J R Soc Interface 12(107):20141405
Thomson M, Gunawardena J (2009) The rational parameterisation theorem for multisite post-translational modification systems. J Theor Biol 261(4):626–636
Tung H-R (2018) Precluding oscillations in Michaelis-Menten approximations of dual-site phosphorylation systems. Math Biosci 306:56–59
Virshup DM, Forger DB (2009) Keeping the beat in the rising heat. Cell 137(4):602–604
Wolfram Research Inc. (2018) Mathematica, Version 11.3, Champaign, IL
Yang X (2002) Generalized form of Hurwitz-Routh criterion and Hopf bifurcation of higher order. Appl Math Lett 15(5):615–621
Acknowledgements
AS was partially supported by the NSF (DMS-1312473/1513364 and DMS-1752672) and the Simons Foundation (#521874). AS thanks Jonathan Tyler for helpful discussions. CC was partially supported by the Deutsche Forschungsgemeinschaft DFG (DFG-284057449).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
A Files in the Supporting Information
A Files in the Supporting Information
The following files can be found as supplementary material:
Text files:
-
mixed_H5N_kb.txt ...contains H5N, the numerator of \(\det H_5\) under the assumption \(k_2=k_6=k_9=k_b\)
-
mixed_W.txt ...contains a matrix W that defines (3)
-
mixed_xt.txt ...contains xt, the parameterization (7)
-
mixed_Jx.txt ...contains Jx, the Jacobian evaluated at the parameterization (7)
Mathematica Notebooks:
-
mixed_analysis_H5N_x1_LT.nb:
Functionality: This file can be used to obtain \(\text {numerator}(\det H_5)\) as in (12), in particular to examine the coefficients \(\alpha _{01}\), \(\alpha _{10}\), ...
Input: the file mixed_H5N_kb.txt
-
mixed_analysis_H5N_x2_LT.nb:
Functionality: This file can be used to obtain \(\text {numerator}(\det H_5)\) as in (14), in particular to examine the coefficients \(\alpha _{0}\), ..., \(\alpha _{3}\) and \(\beta _0\), ..., \(\beta _3\).
Input: the file mixed_H5N_kb.txt
-
mixed_coeffs_charpoly.nb:
Functionality: This file can be used to obtain the characteristic polynomial of the Jacobian of the system (2). It contains the Mathematica commands to establish \(b_i>0\).
Input: the file mixed_Jx.txt
-
mixed_Hi.nb:
Functionality: This file can be used to obtain the determinants of the Hurwitz matrices \(H_2\), ..., \(H_5\). It contains the Mathematica commands to establish \(\det H_i >0\), for \(i=2\), 3, 4 and that \(\det H_5\) is of mixed sign.
Input: the file mixed_Jx.txt
-
mixed_generate_rc.nb:
Functionality: This file contains a realization of Procedure 5.1.
Input: the files mixed_H5N_kb.txt, mixed_W.txt, mixed_xt.txt, mixed_Jx.txt.
Rights and permissions
About this article
Cite this article
Conradi, C., Mincheva, M. & Shiu, A. Emergence of Oscillations in a Mixed-Mechanism Phosphorylation System. Bull Math Biol 81, 1829–1852 (2019). https://doi.org/10.1007/s11538-019-00580-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-019-00580-6